Difference between revisions of "Computational Regulatory Genomics"
>Admin m (Admin moved page Main Page to Computational Regulatory Genomics over redirect) |
|||
| (48 intermediate revisions by 8 users not shown) | |||
| Line 1: | Line 1: | ||
| − | __NOTOC__ | + | <h2>Computational Regulatory Genomics, Gene Regulatory Networks, and Transcriptional Regulation <br> Beer Lab</h2> __NOTOC__ __NOTITLE__ |
| − | __NOTITLE__ | + | <metadesc>Computational Regulatory Genomics, Gene Regulatory Networks, and Transcriptional Regulation in the Beer Lab for graduate PhD and postdoctoral research in BME and IGM at Johns Hopkins, top ranking programs modeling DNA and systems biology.</metadesc> |
| − | <metadesc>Computational Regulatory Genomics for graduate PhD and postdoctoral research in BME and IGM at Johns Hopkins, top ranking programs modeling DNA and systems biology.</metadesc | ||
| − | |||
<h3>[[Recent News ]] [[Lab Members ]] [[Publications]] [[Postdoctoral Positions Available]] [[Resources]]</h3> | <h3>[[Recent News ]] [[Lab Members ]] [[Publications]] [[Postdoctoral Positions Available]] [[Resources]]</h3> | ||
| − | [[File:Beer_Michael_small. | + | [[File:Beer_Michael_small.jpg|link=http://www.bme.jhu.edu/people/faculty/michael-beer/]] [[File:EncodeNatureGraphic_small.png]] [[File:dyn_net.gif]] |
| − | + | [[File:Igvf_logo.png|150px]] [[File:Encode_logo.png]] [[File:Nhgri_logo.jpg|250px]] [[File:hopkins_logo.jpg|200px]] | |
| − | |||
| − | |||
| − | |||
| − | * '''[ | + | We are in the '''[http://www.bme.jhu.edu/people/faculty/michael-beer/ Department of Biomedical Engineering]''' and the '''[https://www.hopkinsmedicine.org/profiles/results/directory/profile/8377361/michael-beer McKusick-Nathans Department of Genetic Medicine]''' at Johns Hopkins University. You can apply for graduate study in my lab through '''[http://www.bme.jhu.edu/people/faculty/michael-beer/ BME]''' or the Ph.D. program in '''[https://www.hopkinsmedicine.org/genetic-medicine/education-training/human-genetics-genomics-graduate-program Human Genetics and Genomics.]''' |
| + | |||
| + | <h3>Research Interests: </h3> The ultimate goal of our research is to understand how gene regulatory information is encoded in genomic DNA sequence. | ||
| + | |||
| + | We have recently made significant progress using AI/ML to understand how DNA sequence specifies cell-type specific enhancer activity, target gene activation through looping constraints, and how dynamic enhancer-gene regulatory networks control cell-fate decisions and contribute to human disease. For details, see: | ||
| + | |||
| + | * '''[https://rdcu.be/dhCoG Dynamic network-guided CRISPRi screen identifies CTCF-loop-constrained nonlinear enhancer gene regulatory activity during cell state transitions.]''' Luo R, et al. Nature Genetics 2023 | ||
| + | |||
| + | * '''[https://genome.cshlp.org/content/34/5/680 Machine learning identifies activation of RUNX/AP-1 as drivers of mesenchymal and fibrotic regulatory programs in gastric cancer.]''' Razavi-Mohseni M, et al. Genome Research 2024 | ||
| + | |||
| + | * '''[https://www.annualreviews.org/doi/abs/10.1146/annurev-genom-121719-010946?journalCode=genom Enhancer Predictions and Genome-Wide Regulatory Circuits.]''' Beer MA, Shigaki D, Huangfu D. Ann. Rev. Genomics and Human Genetics 2020. | ||
| + | |||
| + | * '''[https://onlinelibrary.wiley.com/doi/abs/10.1002/humu.23797 Integration of multiple epigenomic marks improves prediction of variant impact in saturation mutagenesis reporter assay.]''' Shigaki D, et al. Human Mutation 2019. | ||
| + | |||
| + | * '''[https://www.nature.com/articles/ng.3331 A method to predict the impact of regulatory variants from DNA sequence.]''' Lee D, Gorkin DU, Baker M, Strober BJ, Asoni AL, McCallion AS, Beer, MA. Nature Genetics 2015. | ||
* '''[http://www.ploscompbiol.org/article/info%3Adoi%2F10.1371%2Fjournal.pcbi.1003711 Enhanced Regulatory Sequence Prediction Using Gapped k-mer Features.]''' Ghandi M, Lee D, Mohammad-Noori M, and Beer MA. 2014. PLOS Computational Biology. July 17, 2014. | * '''[http://www.ploscompbiol.org/article/info%3Adoi%2F10.1371%2Fjournal.pcbi.1003711 Enhanced Regulatory Sequence Prediction Using Gapped k-mer Features.]''' Ghandi M, Lee D, Mohammad-Noori M, and Beer MA. 2014. PLOS Computational Biology. July 17, 2014. | ||
| − | + | and other [[Publications]]. | |
| − | + | Our work uses functional genomics DNase-seq, ATAC-seq, ChIP-seq, RNA-seq, CRISPR, MPRA, Hi-C, and chromatin state data to computationally identify combinations of transcription factor binding sites which operate to define the activity of cell-type specific enhancers and their activation of target genes. We are currently focused on: | |
| − | * | + | * improving AI/ML methods to build more accurate models of enhancer activity |
| − | * | + | * predicting the impact of mutations on enhancer activity and their contributions to disease |
| − | * | + | * using AI/ML models to build and test dynamic gene regulatory networks with targeted CRISPR perturbation |
| − | + | * using large scale datasets (ENCODE/IGVF) and CRISPR to constrain models of enhancer-promoter interactions and DNA looping mechanisms | |
| − | + | * using this sequence based regulatory code to understand mechanisms of gene regulatory network disruption in cancer and develop therapeutic strategies | |
| − | |||
| − | |||
| − | |||
| − | * using this sequence based regulatory code to | ||
| − | We are located in the McKusick-Nathans Institute of Genetic Medicine, and the Department of Biomedical Engineering, which has long been a leader in the development of rigorous quantitative modeling of biological systems, and is a natural home for graduate studies in Bioinformatics and Computational Biology at Johns Hopkins, including research in | + | We are located in the McKusick-Nathans Institute of Genetic Medicine, and the Department of Biomedical Engineering, which has long been a leader in the development of rigorous quantitative modeling of biological systems, and is a natural home for graduate studies in Genomics, Bioinformatics, and Computational Biology at Johns Hopkins, including research in Systems Biology, AI, Machine Learning, and Dynamic Gene Regulatory Network Modeling. |
| + | <h3>About Computational Biology in JHU Biomedical Engineering:</h3> | ||
| − | |||
The Department of Biomedical Engineering has long been a leader in the development of rigorous quantitative modeling of biological systems, and is a natural home for graduate studies in Bioinformatics and Computational Biology at Johns Hopkins. Students with backgrounds in Physics, Mathematics, Computer Science and Engineering are encouraged to apply. Opportunities for research include: Computational Medicine, Genomics, Systems Biology, Machine Learning, and Network Modeling. Graduate students in Johns Hopkins' Biomedical Engineering programs can select research advisors from throughout Johns Hopkins' Medical Institutions, Whiting School of Engineering, and Krieger School of Arts and Sciences. | The Department of Biomedical Engineering has long been a leader in the development of rigorous quantitative modeling of biological systems, and is a natural home for graduate studies in Bioinformatics and Computational Biology at Johns Hopkins. Students with backgrounds in Physics, Mathematics, Computer Science and Engineering are encouraged to apply. Opportunities for research include: Computational Medicine, Genomics, Systems Biology, Machine Learning, and Network Modeling. Graduate students in Johns Hopkins' Biomedical Engineering programs can select research advisors from throughout Johns Hopkins' Medical Institutions, Whiting School of Engineering, and Krieger School of Arts and Sciences. | ||
| − | <h3>[http://karchinlab.org/bme-compbio-jhu Visit Some Computational Labs at Johns Hopkins]</h3> | + | <!--<h3>[http://karchinlab.org/bme-compbio-jhu Visit Some Computational Labs at Johns Hopkins]</h3> |
| − | <h3>[http://ccb.jhu.edu/ Center for Computational Biology at Johns Hopkins]</h3> | + | <h3>[http://ccb.jhu.edu/ Center for Computational Biology at Johns Hopkins]</h3> --> |
[[File:bmesmall.png]] | [[File:bmesmall.png]] | ||
Latest revision as of 01:24, 31 August 2025
Computational Regulatory Genomics, Gene Regulatory Networks, and Transcriptional Regulation
Beer Lab
Recent News Lab Members Publications Postdoctoral Positions Available Resources
We are in the Department of Biomedical Engineering and the McKusick-Nathans Department of Genetic Medicine at Johns Hopkins University. You can apply for graduate study in my lab through BME or the Ph.D. program in Human Genetics and Genomics.
Research Interests:
The ultimate goal of our research is to understand how gene regulatory information is encoded in genomic DNA sequence.
We have recently made significant progress using AI/ML to understand how DNA sequence specifies cell-type specific enhancer activity, target gene activation through looping constraints, and how dynamic enhancer-gene regulatory networks control cell-fate decisions and contribute to human disease. For details, see:
- Dynamic network-guided CRISPRi screen identifies CTCF-loop-constrained nonlinear enhancer gene regulatory activity during cell state transitions. Luo R, et al. Nature Genetics 2023
- Machine learning identifies activation of RUNX/AP-1 as drivers of mesenchymal and fibrotic regulatory programs in gastric cancer. Razavi-Mohseni M, et al. Genome Research 2024
- Enhancer Predictions and Genome-Wide Regulatory Circuits. Beer MA, Shigaki D, Huangfu D. Ann. Rev. Genomics and Human Genetics 2020.
- Integration of multiple epigenomic marks improves prediction of variant impact in saturation mutagenesis reporter assay. Shigaki D, et al. Human Mutation 2019.
- A method to predict the impact of regulatory variants from DNA sequence. Lee D, Gorkin DU, Baker M, Strober BJ, Asoni AL, McCallion AS, Beer, MA. Nature Genetics 2015.
- Enhanced Regulatory Sequence Prediction Using Gapped k-mer Features. Ghandi M, Lee D, Mohammad-Noori M, and Beer MA. 2014. PLOS Computational Biology. July 17, 2014.
and other Publications.
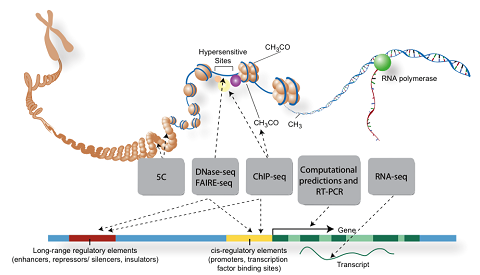
Our work uses functional genomics DNase-seq, ATAC-seq, ChIP-seq, RNA-seq, CRISPR, MPRA, Hi-C, and chromatin state data to computationally identify combinations of transcription factor binding sites which operate to define the activity of cell-type specific enhancers and their activation of target genes. We are currently focused on:
- improving AI/ML methods to build more accurate models of enhancer activity
- predicting the impact of mutations on enhancer activity and their contributions to disease
- using AI/ML models to build and test dynamic gene regulatory networks with targeted CRISPR perturbation
- using large scale datasets (ENCODE/IGVF) and CRISPR to constrain models of enhancer-promoter interactions and DNA looping mechanisms
- using this sequence based regulatory code to understand mechanisms of gene regulatory network disruption in cancer and develop therapeutic strategies
We are located in the McKusick-Nathans Institute of Genetic Medicine, and the Department of Biomedical Engineering, which has long been a leader in the development of rigorous quantitative modeling of biological systems, and is a natural home for graduate studies in Genomics, Bioinformatics, and Computational Biology at Johns Hopkins, including research in Systems Biology, AI, Machine Learning, and Dynamic Gene Regulatory Network Modeling.
About Computational Biology in JHU Biomedical Engineering:
The Department of Biomedical Engineering has long been a leader in the development of rigorous quantitative modeling of biological systems, and is a natural home for graduate studies in Bioinformatics and Computational Biology at Johns Hopkins. Students with backgrounds in Physics, Mathematics, Computer Science and Engineering are encouraged to apply. Opportunities for research include: Computational Medicine, Genomics, Systems Biology, Machine Learning, and Network Modeling. Graduate students in Johns Hopkins' Biomedical Engineering programs can select research advisors from throughout Johns Hopkins' Medical Institutions, Whiting School of Engineering, and Krieger School of Arts and Sciences.